Simulating Multiple Behaviors on Networks
Aníbal Olivera M.
2024-11-21
Source:vignettes/simulating-multiple-behaviors-on-networks.Rmd
simulating-multiple-behaviors-on-networks.RmdNews for netdiffuseR package.
#devtools::install_github("USCCANA/netdiffuseR", ref = "47-split-behaviors-rdiffnet")
library(netdiffuseR)Introduction
- Social networks facilitate the spread of news, gossip, behaviors, and products. Granovetter specified a simple and intuitive mechanism that underpins much of the research on social network diffusion: individuals adopt a behavior if enough others do so.
- More specifically, in an interacting population of individuals where a behavior is spreading, each individual has a particular “threshold” and adopts the behavior if the proportion of others who have already adopted the behavior exceeds the threshold.
- However, adoption booms are commonly followed by periods of bust—riots come to an end, new fashions go out of style, and juicy gossip turns to stale news.
- So the adoption of a fashion is followed by the disadoption of that fashion by a new one.
- This dynamic requires being able to handle more than one ‘propagation’ at a time.
- Until now, netdiffuseR was not able to simulate more than one diffusion in the same setup.
Allowing multi-behavior diffusion simulations
How rdiffnet works until now
In a (very) simplified way, rdiffnet has 4 steps. Before
this current version, the workflow can be diagrammed as:
rdiffnet( graph, seed.p.nodes, seed.nodes, behavior, threshold.dist)
Step 0.0: setting n and t if not provided * this depends on graph
Step 1.0: setting seed nodes * this depends on seed.p.nodes and seed.nodes
Step 2.0: setting threshold for each node * ↳
rdiffnet_make_threshold( threshold.dist, n )
Step 3.0: running the simulation * ↳
exposure( graph, exposure.args = list(...) ) * ↳ allow us
to compute time of adoption (toa)
Step 4.0: creating diffnet object * ↳
new_diffnet( graph, ..., toa, ... )
How rdiffnet works from now
In the current version, we maintain the basic structure of the workflow, but adding some functions and modifying the existing for allowing multi-behavior diffusion.
rdiffnet( graph, seed.p.nodes, seed.nodes, behavior, threshold.dist)
Step 0.0: setting n and t if not provided * this depends on graph
Step 1.0: validate the arguments * ↳
rdiffnet_validate_args(seed.p.adopt, seed.nodes, behavior)* this allows us to set seed nodesStep 2.0: setting threshold for each node * ↳
rdiffnet_make_threshold( threshold.dist, n, num_of_behaviors)Step 3.0: running the simulation * ↳
exposure( graph, exposure.args = list(...) )* ↳disadopt( expo, cumadopt, t)* to compute time of adoption (toa) for each behaviorStep 4.0: creating diffnet object * ↳
new_diffnet( graph, ..., toa, ... )
Step 5.0: splitting behaviors (optional) * ↳
split_behaviors( rdiffnet_multiple_obj )
As you can see, now in Step 1.0, before identify the seed nodes as
initial condition, the inputs seed.p.adopt (default 0.1 for
single behavior), seed.nodes (default ‘random’), and
behavior (default ‘random behavior’) are passed to an
internal function that validates they as accepted inputs and homogenized
the objects for the rest of the code.
As before, rdiffnet still accepts several kinds of
inputs, for different classes classes and specifications, but the input
that characterized the multi-behavior simulation is
exclusively seed.p.adopt. If
class(seed.p.adopt) is a list, then the
simulation will run a total of length(seed.p.adopt)
behaviors in the same setup. This table summarize the possible inputs
for seed.p.adopt, seed.nodes,
threshold.dist, and behavior, showing some
examples.
| Parameter | Type | single | multiple |
|---|---|---|---|
| seed.p.adopt | numeric | 0.1 c(0.1) |
X |
| character | X | X | |
| list | X | list(0.1, 0.05) | |
| seed.nodes | numeric | c(2,4,6) | c(2,4,6) |
| character | “random” c(“random”) |
“random” c(“random”) c(“random”, “central”) |
|
| list | X | list(“random”, “central”) list( c(1,3,5), c(2,4,6) ) |
|
| behavior | numeric | X | X |
| character | “tobacco” c(“tobacco”) |
“tobacco” c(“tobacco”) c(“tobacco”, “alcohol”) |
|
| list | X | list(“tobacco”, “alcohol”) | |
| threshold | numeric | 0.33 rep(0.33, 100) |
0.33 rep(0.33, 100) |
| function | function() runif(1) | function() runif(1) | |
| matrix | X | matrix( runif(100), n_nodes, n_behavior) |
|
| list | X | list( 0.33, 0.66) list( runif(100), runif(100)) list( function() runif(1), function() runif(1)) |
All functions in other steps were revised to allow now different
object that can handle multi-behavior, while still can execute the
already in-build functions for single diffusion within the package (as
plot(), plot_diffnet(), and
plot_adopters(), among many others).
Additionally, now there is a new function
split_behaviors() that returns a list where each element is
a separate rdiffnet object, that correspond with each separate behavior.
So in this way, you could use the same machinery constructed for single
behavior, to plot or analyze now the result of the multi-behavior
simulation.
All those features are shown in more detail below.
rdiffnet() examples
For single diffusion:
## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.09
## Static attributes : real_threshold (1)
## Dynamic attributes : -
?rdiffnet
rdiffnet(100, 5, seed.p.adopt = 0.1)## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.31
## Static attributes : real_threshold (1)
## Dynamic attributes : -
rdiffnet(100, 5, seed.p.adopt = 0.1, behavior = 'tabacco')## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : tabacco
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.27
## Static attributes : real_threshold (1)
## Dynamic attributes : -
rdiffnet(100, 5, seed.p.adopt = 0.1, threshold.dist = 0.3)## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.37
## Static attributes : real_threshold (1)
## Dynamic attributes : -
rdiffnet(100, 5, seed.p.adopt = 0.1, threshold.dist = function(x) 0.3)## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.57
## Static attributes : real_threshold (1)
## Dynamic attributes : -## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.36
## Static attributes : real_threshold (1)
## Dynamic attributes : -
rdiffnet(100, 5, seed.p.adopt = 0.1, seed.nodes = 'central')## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 1.00
## Static attributes : real_threshold (1)
## Dynamic attributes : -## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.48
## Static attributes : real_threshold (1)
## Dynamic attributes : -but also, we can specify the network:
set.seed(121)
n <- 200
t <- 10
graph <- rgraph_ws(n, 10, p=.3) # watts-strogatz model
thr <- runif(n, .3,.5)
rdiffnet(seed.graph = graph, t = t , seed.p.adopt = 0.1, threshold.dist = thr)## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion
## # of nodes : 200 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 10 (1 - 10)
## Type : directed
## Num of behaviors : 1
## Final prevalence : 0.21
## Static attributes : real_threshold (1)
## Dynamic attributes : -For multi diffusion:
## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : tabacco_1, tabacco_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.11, 0.10
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : tabacco, alcohol
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.19, 0.39
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2
## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.
diffnet$vertex.static.attrs## real_threshold.1 real_threshold.2
## 1 0.3 0.3
## 2 0.3 0.3
## 3 0.3 0.3
## 4 0.3 0.3
## 5 0.3 0.3
## 6 0.3 0.3
## 7 0.3 0.3
## 8 0.3 0.3
## 9 0.3 0.3
## 10 0.3 0.3
## 11 0.3 0.3
## 12 0.3 0.3
## 13 0.3 0.3
## 14 0.3 0.3
## 15 0.3 0.3
## 16 0.3 0.3
## 17 0.3 0.3
## 18 0.3 0.3
## 19 0.3 0.3
## 20 0.3 0.3
## 21 0.3 0.3
## 22 0.3 0.3
## 23 0.3 0.3
## 24 0.3 0.3
## 25 0.3 0.3
## 26 0.3 0.3
## 27 0.3 0.3
## 28 0.3 0.3
## 29 0.3 0.3
## 30 0.3 0.3
## 31 0.3 0.3
## 32 0.3 0.3
## 33 0.3 0.3
## 34 0.3 0.3
## 35 0.3 0.3
## 36 0.3 0.3
## 37 0.3 0.3
## 38 0.3 0.3
## 39 0.3 0.3
## 40 0.3 0.3
## 41 0.3 0.3
## 42 0.3 0.3
## 43 0.3 0.3
## 44 0.3 0.3
## 45 0.3 0.3
## 46 0.3 0.3
## 47 0.3 0.3
## 48 0.3 0.3
## 49 0.3 0.3
## 50 0.3 0.3
## 51 0.3 0.3
## 52 0.3 0.3
## 53 0.3 0.3
## 54 0.3 0.3
## 55 0.3 0.3
## 56 0.3 0.3
## 57 0.3 0.3
## 58 0.3 0.3
## 59 0.3 0.3
## 60 0.3 0.3
## 61 0.3 0.3
## 62 0.3 0.3
## 63 0.3 0.3
## 64 0.3 0.3
## 65 0.3 0.3
## 66 0.3 0.3
## 67 0.3 0.3
## 68 0.3 0.3
## 69 0.3 0.3
## 70 0.3 0.3
## 71 0.3 0.3
## 72 0.3 0.3
## 73 0.3 0.3
## 74 0.3 0.3
## 75 0.3 0.3
## 76 0.3 0.3
## 77 0.3 0.3
## 78 0.3 0.3
## 79 0.3 0.3
## 80 0.3 0.3
## 81 0.3 0.3
## 82 0.3 0.3
## 83 0.3 0.3
## 84 0.3 0.3
## 85 0.3 0.3
## 86 0.3 0.3
## 87 0.3 0.3
## 88 0.3 0.3
## 89 0.3 0.3
## 90 0.3 0.3
## 91 0.3 0.3
## 92 0.3 0.3
## 93 0.3 0.3
## 94 0.3 0.3
## 95 0.3 0.3
## 96 0.3 0.3
## 97 0.3 0.3
## 98 0.3 0.3
## 99 0.3 0.3
## 100 0.3 0.3## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.41, 0.27
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.
diffnet$vertex.static.attrs## real_threshold.1 real_threshold.2
## 1 0.3 0.3
## 2 0.3 0.3
## 3 0.3 0.3
## 4 0.3 0.3
## 5 0.3 0.3
## 6 0.3 0.3
## 7 0.3 0.3
## 8 0.3 0.3
## 9 0.3 0.3
## 10 0.3 0.3
## 11 0.3 0.3
## 12 0.3 0.3
## 13 0.3 0.3
## 14 0.3 0.3
## 15 0.3 0.3
## 16 0.3 0.3
## 17 0.3 0.3
## 18 0.3 0.3
## 19 0.3 0.3
## 20 0.3 0.3
## 21 0.3 0.3
## 22 0.3 0.3
## 23 0.3 0.3
## 24 0.3 0.3
## 25 0.3 0.3
## 26 0.3 0.3
## 27 0.3 0.3
## 28 0.3 0.3
## 29 0.3 0.3
## 30 0.3 0.3
## 31 0.3 0.3
## 32 0.3 0.3
## 33 0.3 0.3
## 34 0.3 0.3
## 35 0.3 0.3
## 36 0.3 0.3
## 37 0.3 0.3
## 38 0.3 0.3
## 39 0.3 0.3
## 40 0.3 0.3
## 41 0.3 0.3
## 42 0.3 0.3
## 43 0.3 0.3
## 44 0.3 0.3
## 45 0.3 0.3
## 46 0.3 0.3
## 47 0.3 0.3
## 48 0.3 0.3
## 49 0.3 0.3
## 50 0.3 0.3
## 51 0.3 0.3
## 52 0.3 0.3
## 53 0.3 0.3
## 54 0.3 0.3
## 55 0.3 0.3
## 56 0.3 0.3
## 57 0.3 0.3
## 58 0.3 0.3
## 59 0.3 0.3
## 60 0.3 0.3
## 61 0.3 0.3
## 62 0.3 0.3
## 63 0.3 0.3
## 64 0.3 0.3
## 65 0.3 0.3
## 66 0.3 0.3
## 67 0.3 0.3
## 68 0.3 0.3
## 69 0.3 0.3
## 70 0.3 0.3
## 71 0.3 0.3
## 72 0.3 0.3
## 73 0.3 0.3
## 74 0.3 0.3
## 75 0.3 0.3
## 76 0.3 0.3
## 77 0.3 0.3
## 78 0.3 0.3
## 79 0.3 0.3
## 80 0.3 0.3
## 81 0.3 0.3
## 82 0.3 0.3
## 83 0.3 0.3
## 84 0.3 0.3
## 85 0.3 0.3
## 86 0.3 0.3
## 87 0.3 0.3
## 88 0.3 0.3
## 89 0.3 0.3
## 90 0.3 0.3
## 91 0.3 0.3
## 92 0.3 0.3
## 93 0.3 0.3
## 94 0.3 0.3
## 95 0.3 0.3
## 96 0.3 0.3
## 97 0.3 0.3
## 98 0.3 0.3
## 99 0.3 0.3
## 100 0.3 0.3## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.36, 0.14
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.20, 0.36
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -
rdiffnet(100, 5, seed.p.adopt = list(0.1, 0.08), threshold.dist = list(function(x) 0.3, function(x) 0.2))## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.11, 0.23
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.For example, the first behavior has seed -random-, the second has -central-, etc.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.36, 0.88
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -
set.seed(123)
seed_nodes <- sample(1:100, 10, replace = FALSE)
rdiffnet(100, 5, seed.p.adopt = list(0, 0), seed.nodes = list(seed_nodes, seed_nodes))## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2
## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in rdiffnet(100, 5, seed.p.adopt = list(0, 0), seed.nodes =
## list(seed_nodes, : Set of initial adopters for behavior 1 set to 1.## Warning in rdiffnet(100, 5, seed.p.adopt = list(0, 0), seed.nodes =
## list(seed_nodes, : Set of initial adopters for behavior 2 set to 1.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 100 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 5 (1 - 5)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.29, 0.29
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -Alternatively, we can specify the network:
set.seed(121)
n <- 200
t <- 10
graph <- rgraph_ws(n, t, p=.3) # watts-strogatz model
thr <- runif(n, .3,.5)
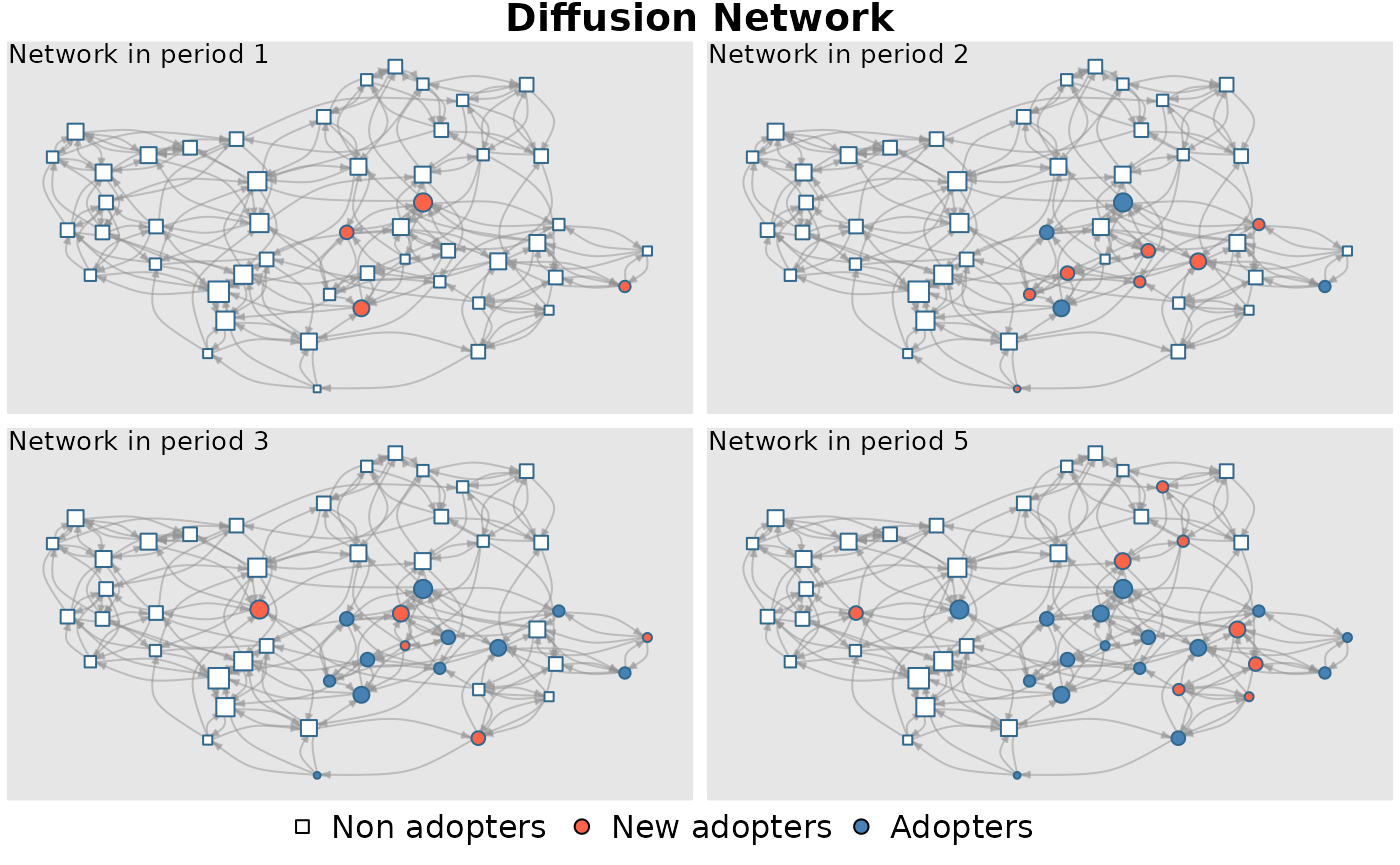
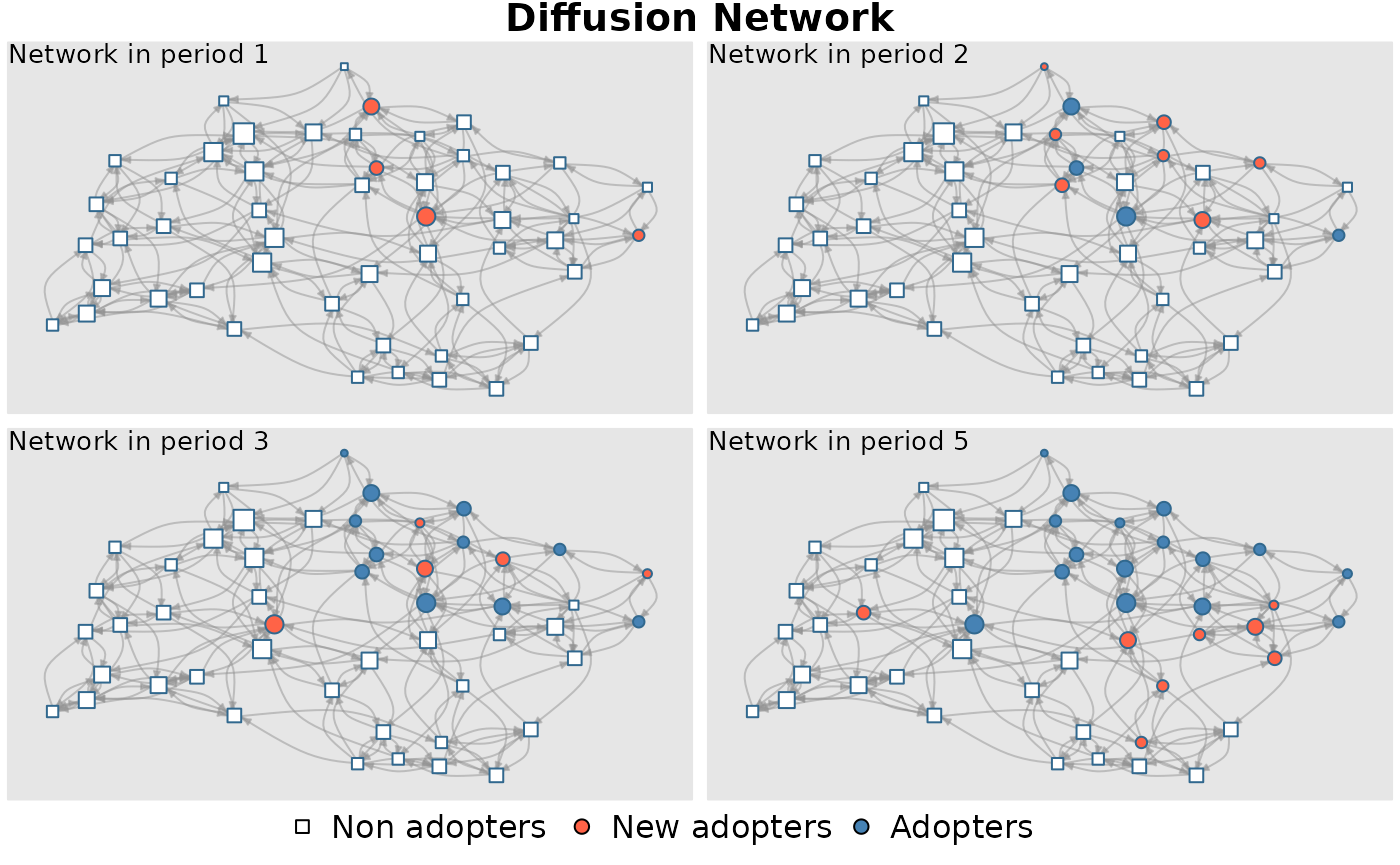
diffnet <- rdiffnet(seed.graph = graph, t = t , seed.p.adopt = list(0.1, 0.15),
threshold.dist = thr)## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.
diffnet## Dynamic network of class -diffnet-
## Name : A diffusion network
## Behavior : Random contagion_1, Random contagion_2
## # of nodes : 200 (1, 2, 3, 4, 5, 6, 7, 8, ...)
## # of time periods : 10 (1 - 10)
## Type : directed
## Num of behaviors : 2
## Prevalence : 0.21, 0.72
## Static attributes : real_threshold.1, real_threshold.2 (2)
## Dynamic attributes : -
split_behaviors() and disadoption
If you want to use other function to analyze the results from the simulation focusing in a single behavior, you could use `split_behaviors()`:
set.seed(12131)
n <- 50
t <- 5
graph <- rgraph_ws(n, 4, p=.3)
seed.nodes <- c(1,5,7,10)
thr <- runif(n, .2,.4)
# Generating identical networks
net_single <- rdiffnet(seed.graph = graph, t = t, seed.nodes = seed.nodes,
seed.p.adopt = 0.1, rewire = FALSE, threshold.dist = thr)
net_multiple <- rdiffnet(seed.graph = graph, t = t, seed.nodes = seed.nodes,
seed.p.adopt = list(0.1, 0.1), rewire = FALSE,
threshold.dist = thr)## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -15710- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.
net_single_from_multiple <- split_behaviors(net_multiple)
net_single_from_multiple_1 <- net_single_from_multiple[[1]]
all(net_single_from_multiple_1$toa == net_single$toa)## [1] NA
all(net_single_from_multiple_1$adopt == net_single$adopt)## [1] TRUE
all(net_single_from_multiple_1$cumadopt == net_single$cumadopt)## [1] TRUEDisadoption
Until now the behaviors are independent, but we can add some
disadoption function to make them dependent each other. This is achieved
by introducing the function you want as an input tho rdiffnet:
rdiffnet <- function(n, t, seed.nodes = "random", seed.p.adopt = 0.05, seed.graph = "scale-free", rgraph.args = list(), rewire = TRUE, rewire.args = list(), threshold.dist = runif(n), exposure.args = list(), name = "A diffusion network", behavior = "Random contagion", stop.no.diff = TRUE, disadopt = NULL)
set.seed(1231)
n <- 500
d_adopt <- function(expo, cumadopt, time) {
# Id double adopters
ids <- which(apply(cumadopt[, time, , drop=FALSE], 1, sum) > 1)
if (length(ids) == 0)
return(list(integer(), integer()))
# Otherwise, make them pick one (literally, you can only adopt
# a single behavior, in this case, we prefer the second)
return(list(ids, integer()))
}
ans_d_adopt <- rdiffnet(n = n, t = 10, disadopt = d_adopt,
seed.p.adopt = list(0.1, 0.1))## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.
tmat <- toa_mat(ans_d_adopt)
should_be_ones_or_zeros <- tmat[[1]]$cumadopt[, 10] + tmat[[2]]$cumadopt[, 10]
all(should_be_ones_or_zeros %in% c(0,1))## [1] TRUE
set.seed(1231)
n <- 100; t <- 5;
graph <- rgraph_ws(n, t, p=.3)
random_dis <- function(expo, cumadopt, time) {
num_of_behaviors <- dim(cumadopt)[3]
list_disadopt <- list()
for (q in 1:num_of_behaviors) {
adopters <- which(cumadopt[, time, q, drop=FALSE] == 1)
if (length(adopters) == 0) {
# only disadopt those behaviors with adopters
list_disadopt[[q]] <- integer()
} else {
# selecting 10% of adopters to disadopt
list_disadopt[[q]] <- sample(adopters, ceiling(0.10 * length(adopters)))
}
}
return(list_disadopt)
}
diffnet_random_dis <- rdiffnet(seed.graph = graph, t = 10, disadopt = random_dis,
seed.p.adopt = list(0.1, 0.1))## Message: Multi-diffusion behavior simulation selected. Number of behaviors: 2## Message: Object -seed.nodes- converted to a -list-.All behaviors will have the same -random- seed nodes.## Message: Name of 1 behavior provided, but 2 are needed. Names generalized to 'behavior'_1, 'behavior'_2, etc.## Warning in new_diffnet(graph = sgraph, toa = toa, self = isself, t0 = 1, :
## Coercing -toa- into integer.
exposure() examples
- Exposure for multiple behaviors:
set.seed(12131)
g <- rgraph_ws(20, 4, p=.3) # watts-strogatz model
set0 <- c(1,5,7,10)
thr <- runif(20, .4,.7)
diffnet <- rdiffnet(seed.graph = g, seed.nodes = set0, t = 4, rewire = FALSE,
threshold.dist = thr)
cumadopt_2 <- diffnet$cumadopt
cumadopt_2 <- array(c(cumadopt_2,cumadopt_2[rev(1:nrow(cumadopt_2)),]), dim=c(dim(cumadopt_2), 2))
print(exposure(diffnet, cumadopt = cumadopt_2))## , , 1
##
## 1 2 3 4
## 1 0.00 0.00 0.25 0.50
## 2 0.25 0.50 0.50 0.75
## 3 0.50 0.50 0.50 0.50
## 4 0.50 0.50 0.75 0.75
## 5 0.00 0.00 0.00 0.25
## 6 0.50 0.75 0.75 0.75
## 7 0.25 0.25 0.50 0.50
## 8 0.00 0.00 0.25 0.75
## 9 0.50 0.50 0.50 0.50
## 10 0.00 0.00 0.25 0.25
## 11 0.00 0.00 0.25 0.50
## 12 0.50 0.50 0.50 0.50
## 13 0.00 0.00 0.00 0.25
## 14 0.00 0.25 0.25 0.25
## 15 0.25 0.25 0.50 0.50
## 16 0.00 0.00 0.00 0.25
## 17 0.25 0.25 0.25 0.50
## 18 0.25 0.25 0.25 0.50
## 19 0.00 0.00 0.00 0.00
## 20 0.25 0.25 0.50 0.50
##
## , , 2
##
## 1 2 3 4
## 1 0.25 0.25 0.50 0.50
## 2 0.25 0.25 0.25 0.50
## 3 0.25 0.50 0.50 0.75
## 4 0.25 0.25 0.25 0.50
## 5 0.00 0.25 0.50 0.50
## 6 0.00 0.00 0.00 0.00
## 7 0.00 0.00 0.00 0.00
## 8 0.25 0.25 0.50 0.75
## 9 0.25 0.25 0.25 0.25
## 10 0.25 0.25 0.25 0.25
## 11 0.00 0.00 0.25 0.25
## 12 0.50 0.50 0.50 0.75
## 13 0.50 0.50 0.75 0.75
## 14 0.25 0.25 0.25 0.25
## 15 0.25 0.50 0.50 0.75
## 16 0.25 0.50 0.75 0.75
## 17 0.00 0.00 0.50 0.75
## 18 0.50 0.50 0.75 0.75
## 19 0.25 0.50 0.50 0.50
## 20 0.25 0.25 0.25 0.50Threshold
Thresholds
- One of the cannonical concepts is the network threshold. Network thresholds (Valente, 1995; 1996), , are defined as the required proportion or number of neighbors that leads you to adopt a particular behavior (innovation), . In (very) general terms
Where is i’s exposure to the innovation and is the adjacency matrix (the network).
split_behaviors()
Examples.
- Importantly, while very useful,
diffnetobjects is not the only way to use netdiffuseR. Most of the functions can also be used with matrices and arrays.
Raw network data
- We call raw network data to datasets that have a somewhat raw form, for example, edgelists,
Note that the echo = FALSE parameter was added to the
code chunk to prevent printing of the R code that generated the
plot.
- For this tutorial, we classify graph data as follows:
- Raw R network data: Datasets with edgelist, attributes, survey data, etc.
- Already R data: already read into R using igraph, statnet, etc.
- Graph files: DL, UCINET, pajek, etc.
- The includes several options to read such data.